- June 2016 (1)
- November 2015 (1)
- September 2015 (1)
- August 2015 (1)
- July 2015 (3)
- June 2015 (3)
- April 2015 (2)
- March 2015 (2)
- December 2014 (1)
- November 2014 (3)
- July 2014 (1)
- May 2014 (1)
- April 2014 (3)
- March 2014 (2)
- February 2014 (4)
- January 2014 (7)
- November 2013 (1)
- October 2013 (2)
- September 2013 (3)
- Informatics (3)
- News (12)
- Request for help - General (10)
- Request for Help - Vote (1)
- Series 4 (16)
On Friday 3rd June 2016, first year students from the Special Studies Program (SSP) at The University of Sydney presented the research performed in their weekly laboratory class. The students held a fantastic film festival, where they premiered shorts (aimed at a HSC audience) about nanoscience. Additionally, students presented research posters based on their synthesis of four brand new antimalarial compounds. We didn't quite get the biological data (i.e. how well the compounds could kill the malaria parasite) in time for the final presentation...but we have them now, so read on!
It's my second year working on a completely open research-focused laboratory course, and once again I've been throughly impressed by our student's research efforts. More coming on the nanoscience project elsewhere, but this blog entry will discuss the SSP lab's contribution to the Open Source Malaria (OSM) consortium and attempt to place their results within the context of the OSM project.
Malaria
Malaria is one of the world’s oldest diseases and remains one of the most deadly. In 2015 there were 213 million recorded cases of malaria and 306 000 deaths, the vast majority being children under the age of five. The current frontline medicines are artemisinin combination therapies (ACTs) that combine a potent antimalarial, artemisinin with other drugs (Figure 1). Artemisinin is isolated from the sweet wormwood plant and was employed in traditional Chinese medicine long before it was marketed as an antimalarial.

Figure 1: Structure of artemisinin, a potent antimalarial
Malaria is a mosquito-borne disease carried by parasites of the genus Plasmodium. There are five strains, of which the most lethal is Plasmodium falciparum.[ii] The parasite is transmitted following a bite from a female anopheles mosquito whilst feeding on blood to nourish her eggs and simultaneously injecting malaria sporozites. The sporozites then travel to the victim’s liver, leading to the development of flu-like symptoms such as high fever and chills.
Although significant progress has been made in the treatment malaria, the emergence of artemisinin resistant strains means that there is an urgent need for a new drug to treat the disease. As malaria mainly affects people in less economically developed countries, there are additional complications associated with disease detection and treatment. Criteria for a new malaria medicine require a single dose cure costing no more than 1 USD. Not only are these criteria difficult to achieve, they also mean that there is a reduced market incentive for large pharmaceutical companies to invest in the development of new therapies.
Open Source Drug Discovery The challenges associated with finding medicines for malaria has led to the development of some more collaborative ways of working to find new drugs. The pharmaceutical company GlaxoSmithKlein (GSK) published the structures of 13 500 compounds (taken from its vast libraries of potential drugs) that were found to be active against malaria.[iii] This was the first time that a pharmaceutical company had shared a large set of data and placed it into the public domain so that researchers all over the world could use the results as starting points for the discovery of new antimalarials. More recently, other companies have followed suit and published the structures of compounds that kill the malaria parasite.[iv] Finding new medicines is a difficult and expensive process; recent statistics have shown that on average it takes 12 years and 2.6 billion USD to bring a drug to market.[v] OSM hope that removing any secrecy from the scientific process might expedite the discovery of drugs and reduce costs by eliminating any unnecessary duplication by researchers in different groups. The other wonderful thing about working in the open is that it elimates many of the barriers to participation, this means that anyone can take part, including the Undergraduate and High School students who have contributed to OSM so far (The University of Sydney, Massachusetts College of Pharmacy and Health Sciences, Lawerence University, Haverford College, The University of Edinburgh, Sydney Grammar School and more to come!)
The Triazolopyrazines
The triazolopyrazines are a group of compounds, first investigated by the pharmaceutical company Pfizer, that were found to kill the malaria parasite. The compounds possess a common ‘core’ (Figure 2, shown in black) which is substituted with different groups of atoms at two positions (Figure 2, shown in blue and green). Medicinal chemists synthesised many molecules of this type and then sent them to biologists who tested their activity. The activity data was used by chemists to design different organic groups to attach to the core in an attempt to optimise the properties of the molecule by increasing activity and enhancing other properties required for a suitable medicine.

Figure 2: The core structure of the triazolopyrazine family of antimalarials.
The first tests performed by biologists were in vitro experiments, meaning that they were tests on cells harvested from their normal environment and observed in glass plates. In theses tests a solution containing a potential new drug is administered to a plate containing Plasmodium falciparum and then biologists measure how effective the molecule is at killing the parasite. These preliminary experiments are really important in determining which molecules are active and inactive, but they don’t provide a full picture of how the molecules will behave in humans. Later experiments on some of the initially promising triazolopyrazine compounds showed that there were some properties that needed to be improved to maximise their chances as a new malaria medicine. Firstly, the solubility of the compounds needed to be increased so that they could be better absorbed in the body. Secondly, the molecules need to be more metabolically stable, so that they are not broken down by the body before they have chance to kill the parasite.
The SSP Project 2015
Over four weeks students worked in small groups (under the guidance of one of four demonstrators) and synthesised a brand new antimalarial compounds following a route developed (and shared online) by OSM.

Figure 3: SSP Synthesis Project
In week one, students were provided with starting material 1 and each demonstrator group was given one of four different aromatic aldehydes (2 in Figure 3, and shown in more detail in Figure 4). A condensation reaction united 1 and 2 to form 3, which was purified by recrystallisation in week 2. In week 3, compound 3 was treated with an oxidant to form bicyclic compound 4 and then in the final week, students displaced the chlorine present in 4 with an alcoholic side chain (Figure 3, shown in blue) to make their target molecules (5).

Figure 4: SSP Aromatic Aldehydes 2015
We combined samples from students who had made the same compounds (each of the four final structures are shown in Figure 5) and purified them in the research lab to make sure that they were clean enough for biolgical testing. Our measurements confirmed that the compounds were pure and so they were vialled up and sent off for testing against the parastie whilst we eagerly awaited the results.

Figure 5: Biological Data SSP 2015
The data showed that phenyl substituted MMV689970 was inactive against the malaria parasite, as were two of the chlorophenyl compounds (MMV689969 and MMV689968). Pleasingly, 4-chlorophenyl substituted MMV663915 was found to be a highly active compund, requiring a very low concentration of the compound to kill the malaria parasite. In fact, MMV663915 turned out to be one of the most active compounds synthesised in the OSM project to date!
The SSP results were warmly welcomed by the OSM team, even the ones that turned out to be dud compounds. In order to develop a model to predict active compounds it's really important to have data for compounds that don't work as well as we might have hoped, as well as the superstars of course! Additionally, one of the best things about being part of an open research project is the fact that we can tell the world which compounds don't work very well, so that no one else wastes time or money working on the same molecules.
Experts from OSM were intrigued by the huge differences in activity between the compounds, particularly between isomers containing a chlorine group. OSM wanted to find out more about which aromatic groups would provide potent antimalarials and wondered whether a substitution in the 4-position (opposite to the bond to the triazolopyrazine core) was of particular importance. This was a job for the SSP Class of 2016!
The SSP Project 2016
In March 2016 a new group of enthusiastic SSP students were ready to build on the work of their 2015 colleagues. A different set of aromatic aldehydes were selected, each substituted with either fluorine or chlorine at the assummed-to-be-important 4-position (Figure 6). Three of the compounds contained an additional fluorine or chlorine in differentt locations around the ring. The students wanted to examine whether it was the presence of a group at position 2 or 3 that had killed the activity in the 2015 compounds, or whether it was the absense of a group at postion 4.

Figure 6: SSP Aromatic Aldehydes 2015
Changing the substituents around the aromatic aldehyde can affect lots of the physical properties of the molecule such as solubility (how greasy vs polar the molecule is) and the molecular weight (the mass of the molecule). It's also important to think about the size of the different substituents, as this can affect how well a molecule is able to fit in a binding site inside a biological system, and can therefore influence how well a molecule can interact with a biological target.

Figure 7: Atomic Radii of Hydrogen, Fluorine and Chlorine
The students successfully made their target molecules and OSM received the biological results for these compounds in June 2016 (Figure 8).

Figure 8 Biological Data SSP 2016
Once again the data was extremely useful. Compounds MMV693151 and MMV693152 were both substituted in the 3- and 4-positions and were found to be only moderately active. It was interesting to compare this result to that for the inactive 3-chloro substituted 2015 compound (MMV689968, Figure 5) as OSM learnt that substitution at the 4-postition is important to maintain activity. Also, as fluorine is similar in size to hydrogen, if we compare MMV689970 with MMV693151 one might infer that the size of the subsituents is less important than other properties that they confer, we see a jump from no activity (MMV689970, Figure 5) to moderate activity (MMV693151, Figure 8). Compound MMV693153, substituted in the 2- and 4-position, was found to be active and fairly good at killing the malaria parasites, again confirming the importance of a substituent at the 4-position. 4-Fluoro substituted MMV693154 supported this theory further as it was also found to be active in the assay.
The SSP Project 2017?
Members of the OSM team across the USA, UK, Australia and NZ are looking at the results obtained as part of the SSP laboratory class and using this data to assist in the design and synthesis of new compounds. OSM particularly want to work on making the active SSP compounds (and others in the project) less greasy, and these analogs will be made and tested in the next few months. The next set of SSP compounds will be designed when all of this information has been taken into full consideration...watch this space!
Get involved!
If you are an undergraduate student or lab coordinator, a PI, a high school teacher or a high school student then OSM would love to hear from you.
As we are an open project, we prefer open communication, so:
Tweet us @O_S_M, find us on G+ or Facebook or come and look at our GitHub thread and see what 'issues' are ongoing in the project.
You can also check out our landing page opensourcemalaria.org and if you would really, really prefer to email then that's ok too: [email protected].
References
WHO World Malaria Report 2015
[iii] Gamo F-J.; et al. Nature. 2010, 465, 305
[v] D. W. Light and J. R. Lexchin, British Medical Journal 2012, 345, e4348. (DOI: 10.1136/bmj.e4348)
Planning for the next round of synthesis is taking place over on GHI358.
This post only contains the strings for all the molecules mentioned there.
Nemesis OC(C1=CC=CC=C1)COC2=CN=CC3=NN=C(C4CNCCC4)N32
InChI=1S/C18H21N5O2/c24-15(13-5-2-1-3-6-13)12-25-17-11-20-10-16-21-22-18(23(16)17)14-7-4-8-19-9-14/h1-3,5-6,10-11,14-15,19,24H,4,7-9,12H2
SNDLBGDEOQPBOQ-UHFFFAOYSA-N
CRO OC(C1=CC=CC=C1)COC2=CN=CC3=NN=C(C4COC4)N32 InChI=1S/C16H16N4O3/c21-13(11-4-2-1-3-5-11)10-23-15-7-17-6-14-18-19-16(20(14)15)12-8-22-9-12/h1-7,12-13,21H,8-10H2 ZLHKTQHFEDMDIZ-UHFFFAOYSA-N
NW Aliphatic FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(OCC(C)(C)OC)N32 InChI=1S/C17H18F2N4O3/c1-17(2,24-3)10-25-14-9-20-8-13-21-22-15(23(13)14)11-4-6-12(7-5-11)26-16(18)19/h4-9,16H,10H2,1-3H3 QPGDQEMNKJULOO-UHFFFAOYSA-N
Sydney Grammar Pyridine OC(C1=CC=CC=C1)COC2=CN=CC3=NN=C(C4=CN=CC=C4)N32 InChI=1S/C18H15N5O2/c24-15(13-5-2-1-3-6-13)12-25-17-11-20-10-16-21-22-18(23(16)17)14-7-4-8-19-9-14/h1-11,15,24H,12H2 GXHFRJHPAQDHIG-UHFFFAOYSA-N
Dimethyl Amine FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(OCC(C4=CC=CC=C4)N(C)C)N32 InChI=1S/C22H21F2N5O2/c1-28(2)18(15-6-4-3-5-7-15)14-30-20-13-25-12-19-26-27-21(29(19)20)16-8-10-17(11-9-16)31-22(23)24/h3-13,18,22H,14H2,1-2H3 WSXRPWJACRAQOI-UHFFFAOYSA-N
Homologous Alcohol FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(OCC(C4=CC=CC=C4)CO)N32 InChI=1S/C21H18F2N4O3/c22-21(23)30-17-8-6-15(7-9-17)20-26-25-18-10-24-11-19(27(18)20)29-13-16(12-28)14-4-2-1-3-5-14/h1-11,16,21,28H,12-13H2 MGANJQKOPZQELF-UHFFFAOYSA-N
Homologous Amine FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(OCC(C4=CC=CC=C4)CN)N32 InChI=1S/C21H19F2N5O2/c22-21(23)30-17-8-6-15(7-9-17)20-27-26-18-11-25-12-19(28(18)20)29-13-16(10-24)14-4-2-1-3-5-14/h1-9,11-12,16,21H,10,13,24H2 ZAYFESUSYJEUPU-UHFFFAOYSA-N
Shackleford FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(OC(C)C(C4=CC=CC=C4)O)N32 InChI=1S/C21H18F2N4O3/c1-13(19(28)14-5-3-2-4-6-14)29-18-12-24-11-17-25-26-20(27(17)18)15-7-9-16(10-8-15)30-21(22)23/h2-13,19,21,28H,1H3 XGFALCXCRQYLSD-UHFFFAOYSA-N
Swain FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(OC(C)C4=CC=C(Cl)C(Cl)=C4)N32 InChI=1S/C20H14Cl2F2N4O2/c1-11(13-4-7-15(21)16(22)8-13)29-18-10-25-9-17-26-27-19(28(17)18)12-2-5-14(6-3-12)30-20(23)24/h2-11,20H,1H3 ILDZQTSEZGQWFR-UHFFFAOYSA-N
Reversed Amide FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(NC(C4=CC=NC(C(F)(F)F)=C4)=O)N32 InChI=1S/C19H11F5N6O2/c20-18(21)32-12-3-1-10(2-4-12)16-29-28-15-9-25-8-14(30(15)16)27-17(31)11-5-6-26-13(7-11)19(22,23)24/h1-9,18H,(H,27,31) NMZQYYGTOGZVCK-UHFFFAOYSA-N
MMV669848 FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(CN4CC(C=CC=C5)=C5C4)N32 InChI=1S/C21H17F2N5O/c22-21(23)29-18-7-5-14(6-8-18)20-26-25-19-10-24-9-17(28(19)20)13-27-11-15-3-1-2-4-16(15)12-27/h1-10,21H,11-13H2
MMV668957 FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(N4CCC(C5=CC=CC=C5)C4)N32 InChI=1S/C22H19F2N5O/c23-22(24)30-18-8-6-16(7-9-18)21-27-26-19-12-25-13-20(29(19)21)28-11-10-17(14-28)15-4-2-1-3-5-15/h1-9,12-13,17,22H,10-11,14H2
MMV669304 FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(CCCC4=CC=CC=C4)N32 InChI=1S/C21H18F2N4O/c22-21(23)28-18-11-9-16(10-12-18)20-26-25-19-14-24-13-17(27(19)20)8-4-7-15-5-2-1-3-6-15/h1-3,5-6,9-14,21H,4,7-8H2
MMV671652 FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(N[C@H](C4=CC=C(F)C(F)=C4)CO)N32 InChI=1S/C20H15F4N5O2/c21-14-6-3-12(7-15(14)22)16(10-30)26-17-8-25-9-18-27-28-19(29(17)18)11-1-4-13(5-2-11)31-20(23)24/h1-9,16,20,26,30H,10H2/t16-/m0/s1
MMV669008 FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(N4CC(OC5=CC(F)=C(F)C=C5)C4)N32 InChI=1S/C21H15F4N5O2/c22-16-6-5-14(7-17(16)23)31-15-10-29(11-15)19-9-26-8-18-27-28-20(30(18)19)12-1-3-13(4-2-12)32-21(24)25/h1-9,15,21H,10-11H2
MMV669353 FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(N4CC(C5=CC=CC=C5)C4)N32 InChI=1S/C21H17F2N5O/c22-21(23)29-17-8-6-15(7-9-17)20-26-25-18-10-24-11-19(28(18)20)27-12-16(13-27)14-4-2-1-3-5-14/h1-11,16,21H,12-13H2
MMV671653 FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(N4N=CC(C5=CC(F)=C(F)C=C5)=C4)N32 InChI=1S/C21H12F4N6O/c22-16-6-3-13(7-17(16)23)14-8-27-30(11-14)19-10-26-9-18-28-29-20(31(18)19)12-1-4-15(5-2-12)32-21(24)25/h1-11,21H
MMV671678 FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(N4C=CN=C4CC5=CC(F)=C(F)C=C5)N32 InChI=1S/C22H14F4N6O/c23-16-6-1-13(9-17(16)24)10-18-28-7-8-31(18)20-12-27-11-19-29-30-21(32(19)20)14-2-4-15(5-3-14)33-22(25)26/h1-9,11-12,22H,10H2
MMV669360 FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(COCC4=CC=C(F)C(F)=C4)N32 InChI=1S/C20H14F4N4O2/c21-16-6-1-12(7-17(16)22)10-29-11-14-8-25-9-18-26-27-19(28(14)18)13-2-4-15(5-3-13)30-20(23)24/h1-9,20H,10-11H2
MMV670243 FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(CCCC4=CC(F)=C(F)C=C4)N32 InChI=1S/C21H16F4N4O/c22-17-9-4-13(10-18(17)23)2-1-3-15-11-26-12-19-27-28-20(29(15)19)14-5-7-16(8-6-14)30-21(24)25/h4-12,21H,1-3H2
The Crazy Hemiacetal FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(OC(O)C(O)C4=CC=CC=C4)N32 InChI=1S/C20H16F2N4O4/c21-20(22)29-14-8-6-13(7-9-14)18-25-24-15-10-23-11-16(26(15)18)30-19(28)17(27)12-4-2-1-3-5-12/h1-11,17,19-20,27-28H LBNIPJYOUJPJNQ-UHFFFAOYSA-N
The Ketone FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(C(CC4=CC=NC(C(F)(F)F)=C4)=O)N32 InChI=1S/C20H12F5N5O2/c21-19(22)32-13-3-1-12(2-4-13)18-29-28-17-10-26-9-14(30(17)18)15(31)7-11-5-6-27-16(8-11)20(23,24)25/h1-6,8-10,19H,7H2 NRTCUOWGNGMJIN-UHFFFAOYSA-N
The Sulfoxide FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C([S+]([O-])CC4=CC=NC(C(F)(F)F)=C4)N32 InChI=1S/C19H12F5N5O2S/c20-18(21)31-13-3-1-12(2-4-13)17-28-27-15-8-25-9-16(29(15)17)32(30)10-11-5-6-26-14(7-11)19(22,23)24/h1-9,18H,10H2 JITAVHCKEWUUSI-UHFFFAOYSA-N
The Sulfone FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(S(CC4=CC=NC(C(F)(F)F)=C4)(=O)=O)N32 InChI=1S/C19H12F5N5O3S/c20-18(21)32-13-3-1-12(2-4-13)17-28-27-15-8-25-9-16(29(15)17)33(30,31)10-11-5-6-26-14(7-11)19(22,23)24/h1-9,18H,10H2 DHPQHOFGWSYVQF-UHFFFAOYSA-N
The Urea FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(N([H])C(N([H])C4=CC(C(F)(F)F)=NC=C4)=O)N32 InChI=1S/C19H12F5N7O2/c20-17(21)33-12-3-1-10(2-4-12)16-30-29-15-9-25-8-14(31(15)16)28-18(32)27-11-5-6-26-13(7-11)19(22,23)24/h1-9,17H,(H2,26,27,28,32) MXAOEFDAAVOYKX-UHFFFAOYSA-N
The Triazole FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(C4=CN(C5=CC(C(F)(F)F)=NC=C5)[N][N]4)N32 InChI=1S/C20H11F5N8O/c21-19(22)34-13-3-1-11(2-4-13)18-30-29-17-9-26-8-15(33(17)18)14-10-32(31-28-14)12-5-6-27-16(7-12)20(23,24)25/h1-10,19H XXNFDIRYVMBYOF-UHFFFAOYSA-N
The Burns FC(F)OC(C=C1)=CC=C1C2=NN=C3C=NC=C(C(C(F)(F)F)N([H])C4=CC=NC(C(F)(F)F)=C4)N32 InChI=1S/C20H12F8N6O/c21-18(22)35-12-3-1-10(2-4-12)17-33-32-15-9-29-8-13(34(15)17)16(20(26,27)28)31-11-5-6-30-14(7-11)19(23,24)25/h1-9,16,18H,(H,30,31) ORWCWEGIHLHCSF-UHFFFAOYSA-N
The Tianyi OC(C1=CC=CC=C1)COC2=CN=CC3=NC(C)=C(C4=CC=C(OC(F)F)C=C4)N32 InChI=1S/C22H19F2N3O3/c1-14-21(16-7-9-17(10-8-16)30-22(23)24)27-19(26-14)11-25-12-20(27)29-13-18(28)15-5-3-2-4-6-15/h2-12,18,22,28H,13H2,1H3 VGNPLLBRBOUQAA-UHFFFAOYSA-N
Chase Hop1 FC(C=C1)=C(F)C=C1CCOC2=CN=CC(N3N=NN=C3C4=CC=C(Cl)C=C4)=N2 InChI=1S/C19H13ClF2N6O/c20-14-4-2-13(3-5-14)19-25-26-27-28(19)17-10-23-11-18(24-17)29-8-7-12-1-6-15(21)16(22)9-12/h1-6,9-11H,7-8H2 HHZUPBCYYAXEQO-UHFFFAOYSA-N
Chase Hop2 FC(C=C1)=C(F)C=C1CCOC2=NC=NC(NC3=NN=C(C4=CC=C(Cl)C=C4)S3)=N2 InChI=1S/C19H13ClF2N6OS/c20-13-4-2-12(3-5-13)16-27-28-19(30-16)26-17-23-10-24-18(25-17)29-8-7-11-1-6-14(21)15(22)9-11/h1-6,9-10H,7-8H2,(H,23,24,25,26,28) BDGBDNWYLOQTMB-UHFFFAOYSA-N
MMV639565 OSM-S-272 FC1=C(F)C=CC(CCOC2=CN=CC3=NN=C(C4=CC=C(Cl)C=C4)N32)=C1 InChI=1S/C19H13ClF2N4O/c20-14-4-2-13(3-5-14)19-25-24-17-10-23-11-18(26(17)19)27-8-7-12-1-6-15(21)16(22)9-12/h1-6,9-11H,7-8H2 PMHUSEXABGDNGS-UHFFFAOYSA-N
MMV669846 OSM-S-273 FC1=C(F)C=CC(CCOC2=CN=CC3=NC=C(C4=CC=C(Cl)C=C4)N32)=C1 InChI=1S/C20H14ClF2N3O/c21-15-4-2-14(3-5-15)18-10-25-19-11-24-12-20(26(18)19)27-8-7-13-1-6-16(22)17(23)9-13/h1-6,9-12H,7-8H2 MQHQNFQVEXYAMO-UHFFFAOYSA-N
MMV670250 OSM-S-274 FC1=C(F)C=CC(CCOC2=CN=CC3=CN=C(C4=CC=C(Cl)C=C4)N32)=C1 InChI=1S/C20H14ClF2N3O/c21-15-4-2-14(3-5-15)20-25-11-16-10-24-12-19(26(16)20)27-8-7-13-1-6-17(22)18(23)9-13/h1-6,9-12H,7-8H2 JHORUMGXLQTHAD-UHFFFAOYSA-N
This post authored by Mat Todd.
At the end of July OSM announced our graphical abstract competition. We wanted a striking single image (that summed up the science and open methods employed in our work on series one) to accompany our upcoming publication.
The competition called for open submissions and we were delighted to receive two wonderful entries, which the judges (Justine Alltimes, John Overington, Javier Gamo, Mat Todd and Alice Williamson) struggled to choose between. Here goes...
First Prize
The judges felt that Viputheshwar Sitaraman's image displayed the science and open methodology in an extremely clear and attractive manner and effectively conveyed the essence of the paper in a single image.

Viputheshwar Sitaraman's image will be published as the graphical abstract for our next paper and he also wins $600 - Congratulations! Note that following the end of the competition, the winning entry was tweaked slightly to make it acceptable for submission to a journal alongside the paper. This version is attached (Revised Final Winner.psd and .jpg). Please use this version when discussing OSM.
Second Prize
The judges also loved the beautiful entry by Merinda Jayne Ramage. This stunning image really illustrated the citizen science aspect of the project and the judges thought she captured the teamwork behind the science. OSM really look forward to using Merinda's image on some upcoming projects. Merinda wins $200.

A HUGE thank you to both Viputheshwar and Merinda - OSM were thrilled to receive work from two talented artists and we hope that this project is the first of many collaborations with artists.
Licence: These images may be used with CC-BY attribution to this page and the name of the original artist.
Post orginally Authored by Alice Williamson
In the past two months, I was doing Open Source Malaria project in lab with Alice Williamson. My project came to an end as the end of my exchange was approaching. The aim of my project was evaluating the possibility of using NBS bromination and cross-coupling reaction to synthesize Series 4 Triazolopyrazine Series Compounds. The importance of this exploration is that it enables the more late-stage diversification of the triazolopyrazine core structure, which is more promising than the current approach. (Picture below shows current approach(top) and new approach(bottom))
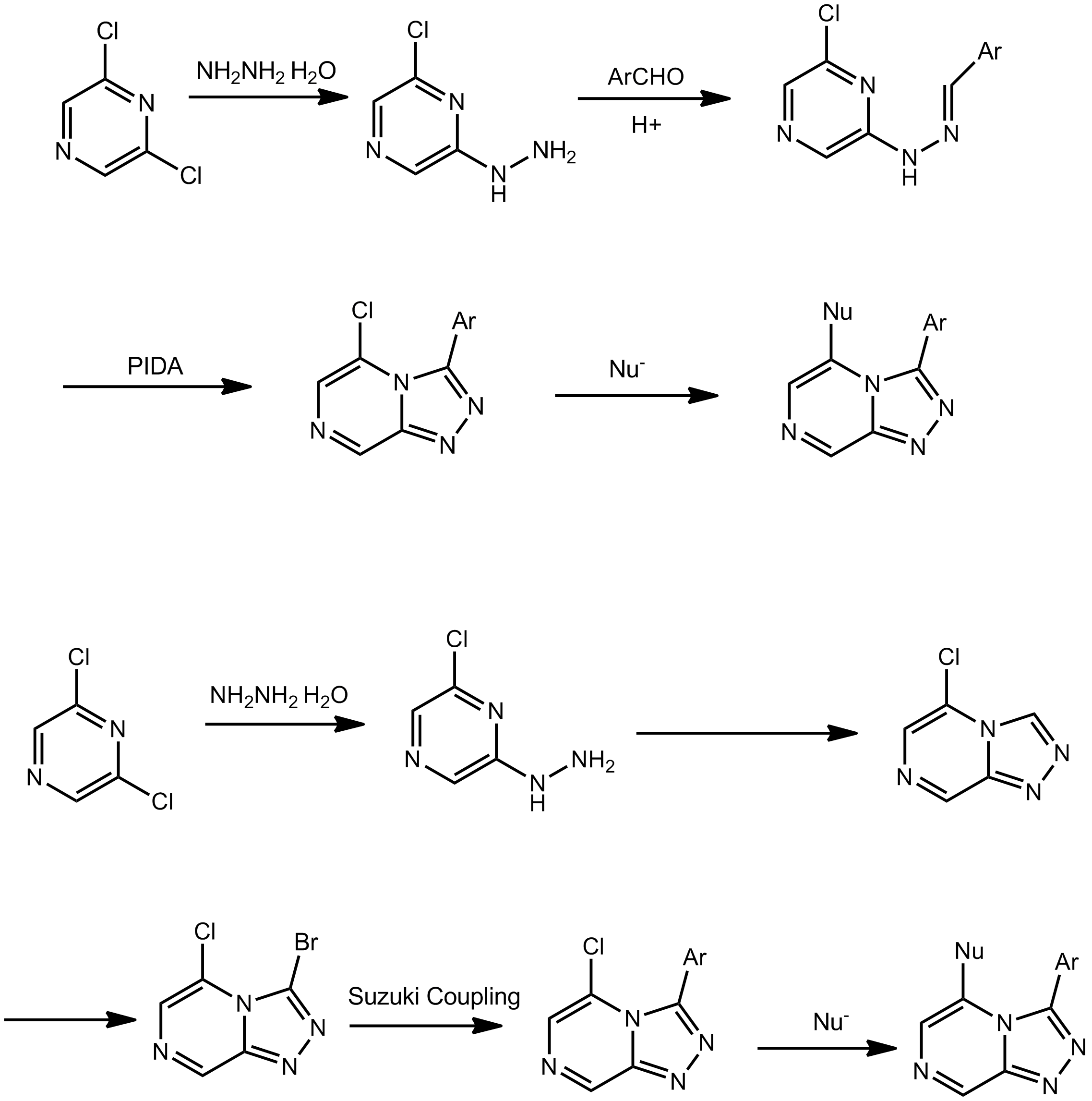
Here is a summary of what I’ve done and accomplished during my stay.
With the preliminary results of this area in hand, we firstly repeated the known reactions to make the triazolopyrazine. H NMR confirmed the right structure. We started the NBS bromination reaction straight after that. According to H NMR, we’ve known that the desired mono-brominated product was successfully obtained. The remaining problem was that the product was hard to be purified and isolated. We’ve tried recrystallization and chromatography to get the relatively pure product with a bad yield. The reasons for this problem would probably be a) generation of multi-brominated byproducts b) remaining staring materials which was not brominated by NBS c) residue of NBS and its byproducts d) similar polarity between the products with bromine in different positions. To optimize the reaction, we used acetic acid as catalyst to facilitate the reaction. Comparison of H NMR revealed a possibility of switched selectivity due to the presence of acetic acid which was exciting, although there was no positive sign showing that the reaction was facilitated.
Suzuki coupling reaction was another focus of my project. With the relative pure products obtained in bromination reaction, we started the Suzuki reaction with different conditions and obtained the arylated products according to H NMR results. What surprised us was there might be a switch of arylation selectivity in the reaction, which was interesting. Although we did not optimize the reaction systematically, we still got some valuable clues in these attempts which enabled a broader exploration on reaction conditions.
Another part of my work was to develop new method to synthesize triazolopyrazine structure. We found an alternative way to make the compound where no further purification was required. But the yield was not satisfactory.
Apart from the research work in the project, I also enjoyed the process where I was able to improve my ability to search literature and write project description. More information about bromination reaction was included in the description paper. Also, I want to thank the group for giving me an opportunity to elaborate my project in front of all members. It was an enjoyable thing which improved my ability to explain academic topic to others clearly and systematically.
I really enjoyed the past eight weeks I spent with Mat, Alice and other group members, had great research and life experience. It was a great honor for me to work with so many nice and excellent people who helped me out during my stay. I would recommend this project to other students in Nanjing University who are interested in this exchange program, spread the open science idea to others and encourage more people to contribute to this project. I would like to thank Prof. Mat Todd for providing me with this stage to do some research and his instructions, thank Alice Williamson for all the care and instructions not only in academic work but also in daily life. Thanks again for the good memory you gave me.
All the best,
Tianyi Zheng (exchange student from Nanjing University)
This past month I've had the opportunity to do some work in the lab with Alice Williamson. My aim was the workup of four compounds (SSP-1:4) synthesised in an undergraduate lab
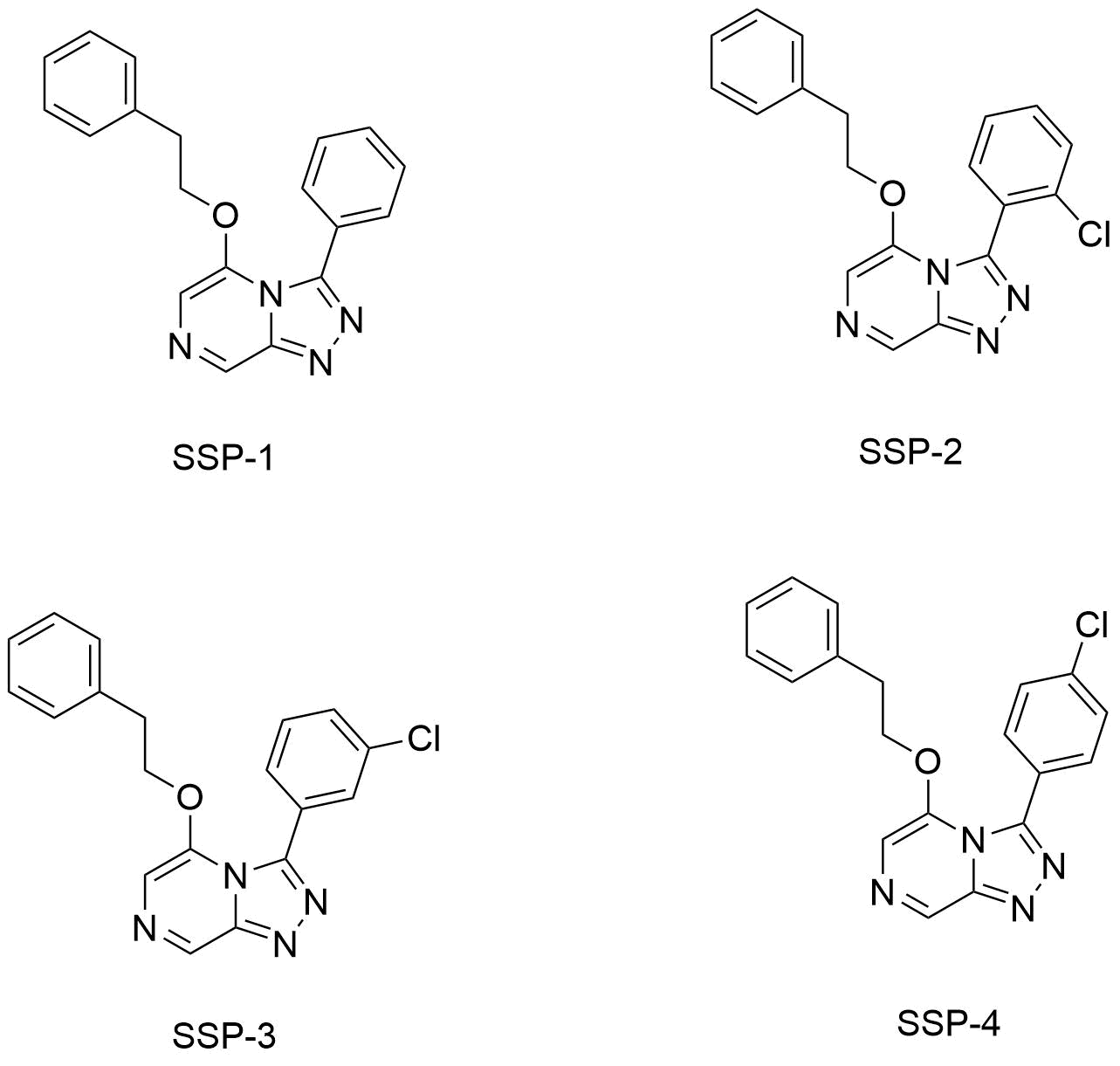
here's a short summary of what I was able to accomplish:
SSP-1: Once the bulk of the original solvent was removed, we ran SSP-1 through a silica column. However 1H NMR data suggested that there was still some impurities in the product. We thus ran the product through a second column. 1H NMR data looks promising, but still under analysis.
SSP-2: SSP-2 was succesfully worked up, with promising 1H NMR data.
SSP-3: SSP-3 was also succesfully worked up with promising 1H NMR data.
SSP-4: Unfortunately due to time constraints I wasn't able to finish off the work up of SSP-4, I presume that it will be finished up by Alice however.
Overall I really enjoyed my time in the lab, met some great people and just had a great experience. I would recommend this to anyone, especially to those doing undergraduate studies in related Science degrees.I would like to thank again Alice Williamson and Mat Todd for providing me with the opportunity and taking the time to help me out on the way.
All the best,
Sebastien Dath